In The News
- USA Today
Withstanding hurricanes, lightning strikes, pests: 'This tree is a survivor'
- USA Today
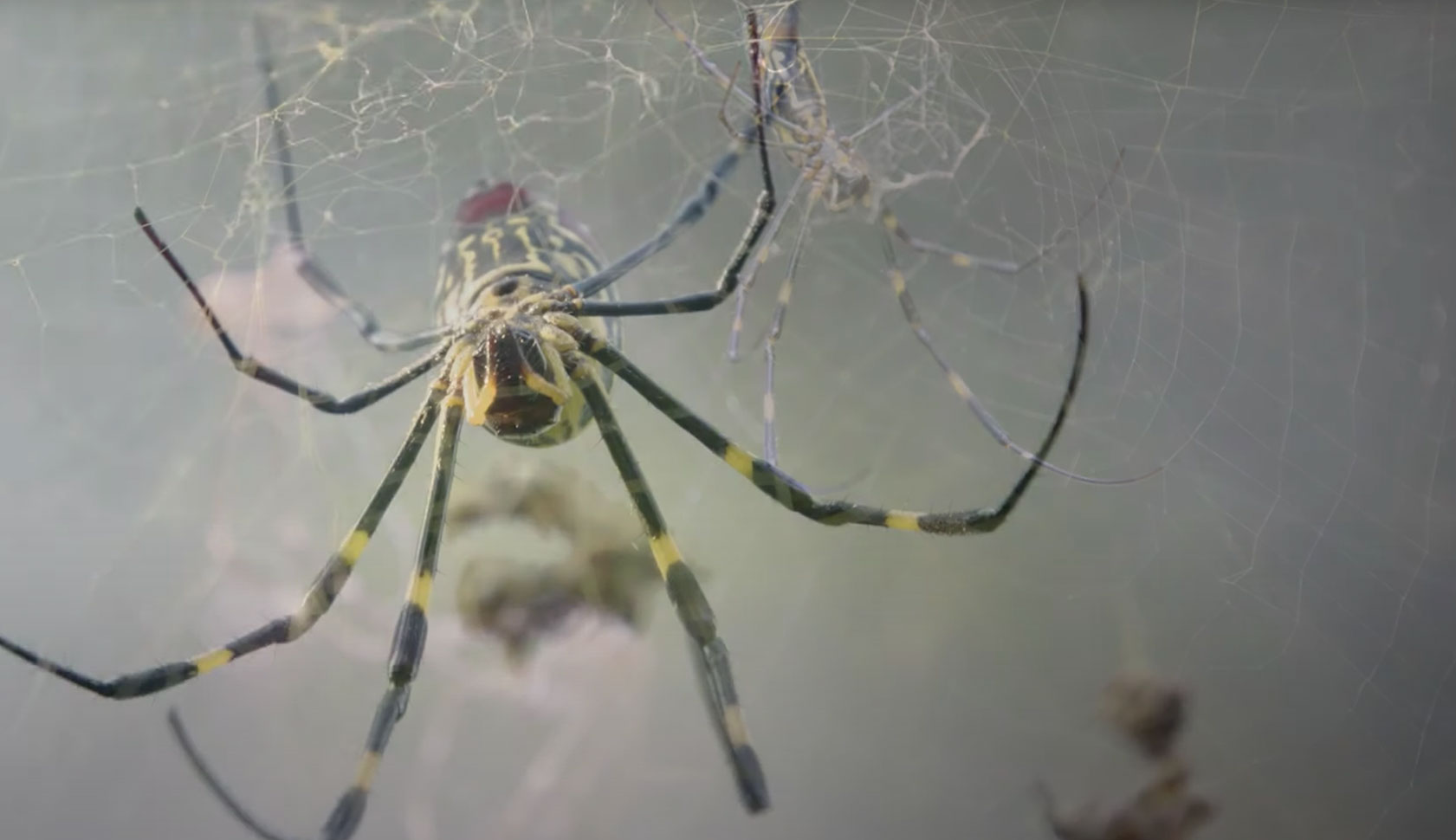
Murder hornet's yellow-legged relative found in Georgia
- AJC
Embrace the buzz: What you need to know about the 2024 cicada invasion
- Georgia Public Broadcasting
Georgia archaeological sites in danger due to sea level rise
- Bloomberg
Black-owned autonomous grocer goes where other stores aren't
In The News
- USA Today
Withstanding hurricanes, lightning strikes, pests: 'This tree is a survivor'
- USA Today
Murder hornet's yellow-legged relative found in Georgia
- AJC
Embrace the buzz: What you need to know about the 2024 cicada invasion
- Georgia Public Broadcasting
Georgia archaeological sites in danger due to sea level rise
- Bloomberg
Black-owned autonomous grocer goes where other stores aren't
ECOLOGY
July 20, 2021
March 22, 2022
Most Popular Features
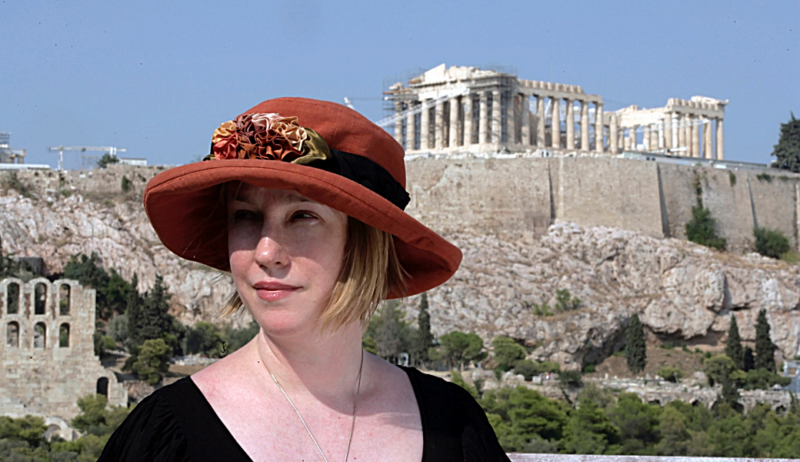
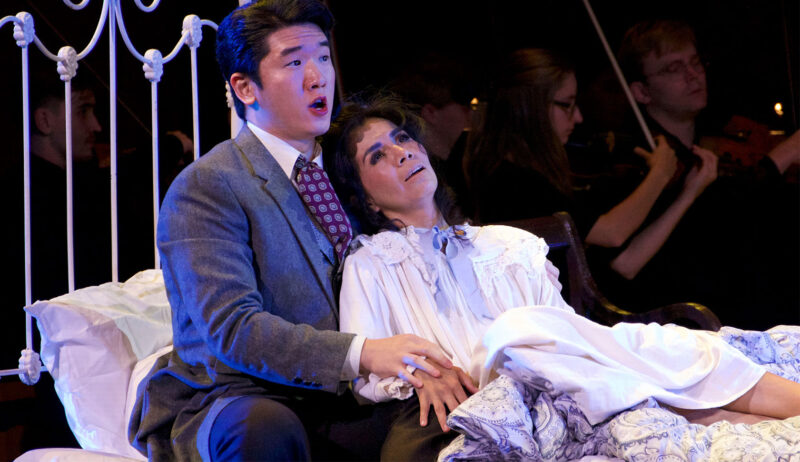
HUMANITIES & ARTS
February 26, 2024
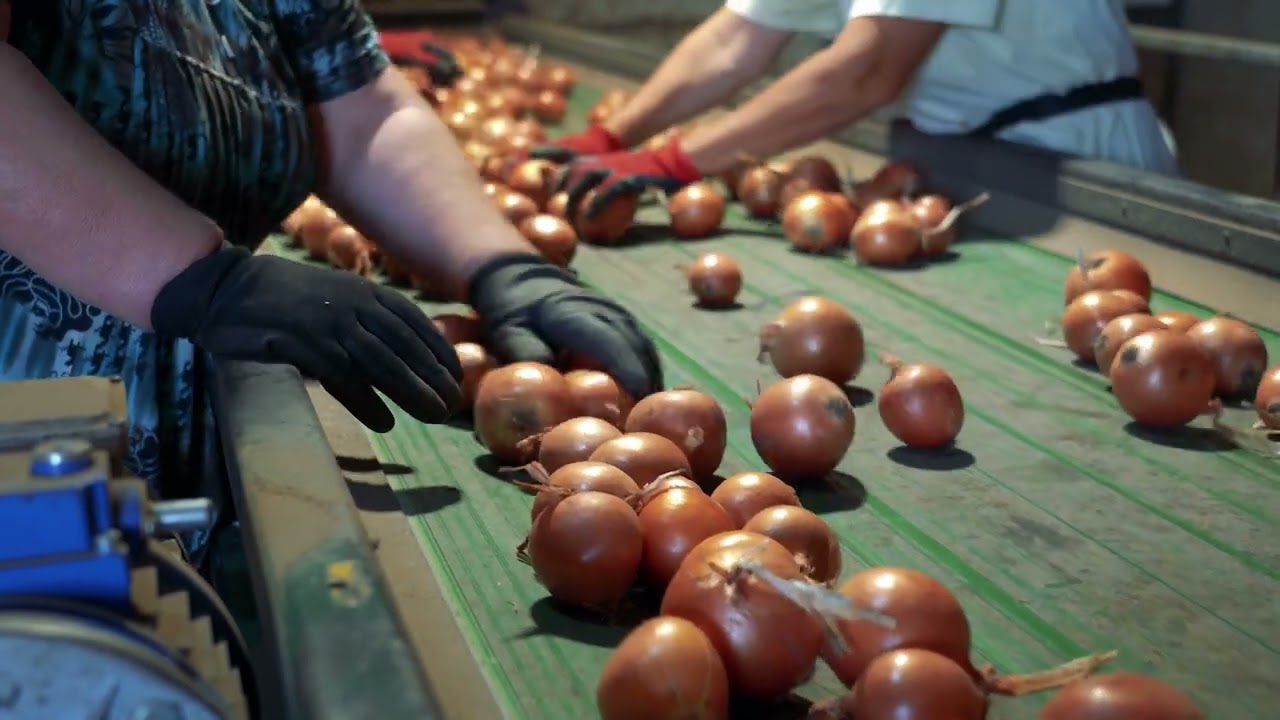
November 30, 2023
Watch
Stay Connected
UGA Research Newsletter
Delivered to your inbox once every month.
Look
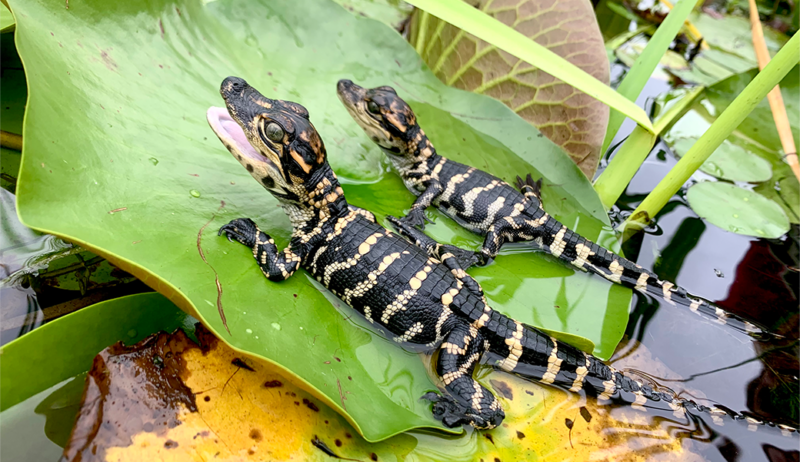
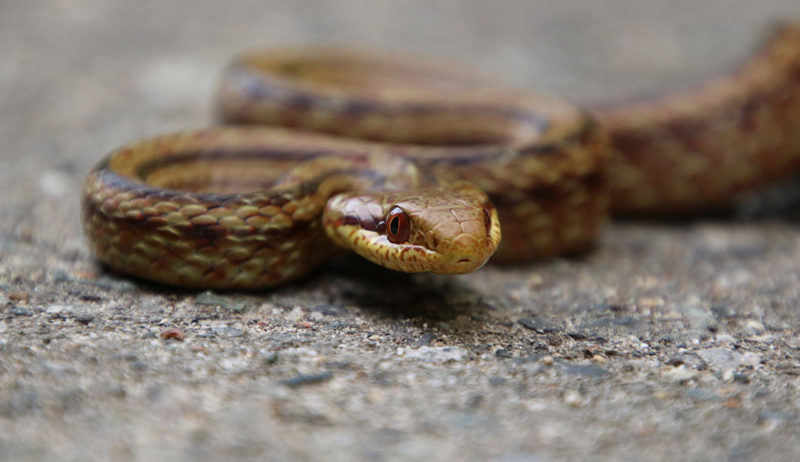
Wildlife
March 7, 2024